Perlin Lab Research
Dr. Perlin with students and colleagues |
EVOLUTION OF HOST/PATHOGEN INTERACTIONS: MECHANISTIC INSIGHTS
The research in my lab focuses on the evolution of interactions between pathogens and the hosts on which they cause disease. At present, this work has two main areas of emphasis: fungal/plant interactions and population dynamics of bacteria resistant to antibiotics.
Contents:
Fungal/Plant Interactions leading to Pathogenesis
Our work with fungal/plant interactions has examined primarily the “smut” fungi.
Plant Diseases Caused by Smut Fungi:
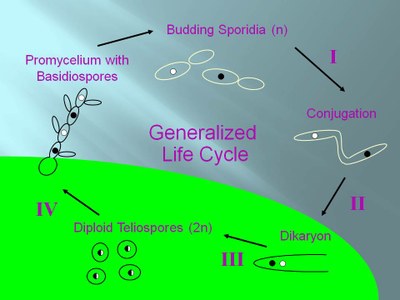
These are basidiomycetous fungi that have several different morphological stages in the course of their development. The final stage is obligately parasitic, in that production of the diploid spores requires successful infection of a suitable host plant. The general lifecycle is illustrated below.
|
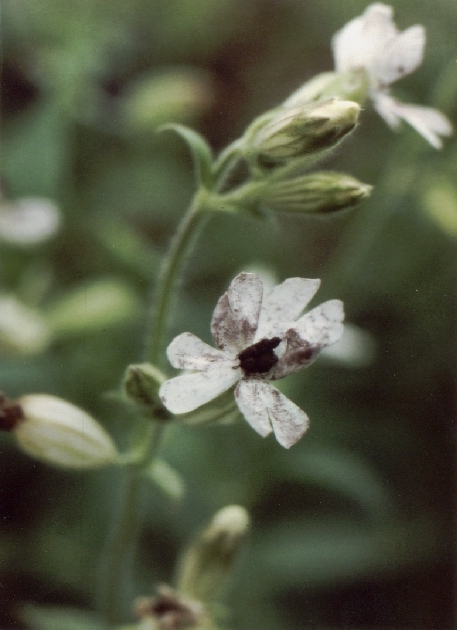
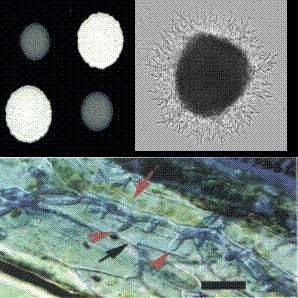
Microbotryum violaceum "anther smut" |
and
|
|
Ustilago maydis "corn smut" |
|
Representative life cycle of a smut fungus |
Questions of Host Specificity and Evolution of Sexual Systems
We began our work with smut fungi over 20 years ago with the anther smut, Microbotryum violaceum, a fungus that infects members of the carnation family (Caryophyllaceae) and replaces the pollen of anthers with its own teliospores. Our early work attempted to establish such molecular methods as genetic transformation and provided DNA “fingerprints” for individual isolates from different host species. Since then it has become clear that what has been called Microbotryum violaceum (or Ustilagoviolacea, Pers. Roussel ) for hundreds of years is actually a species complex, comprised of many different species or incipient species, each of which is adapted to infection of its own host species. What has emerged is an exciting system for examination of the ecology and evolution of host/pathogen interactions in “wild” non-agricultural environments, a model for studying the evolution of sexual systems and sex chromosomes, and gene expression during infection that may give clues to important genes involved in pathogenicity. http://www.amherst.edu/~mhood/mv/community.htm
Fungal Dimorphism and the Link to Pathogenicity
A major aspect of our work with fungi involves genes that control morphogenesis and pathogenicity in fungi. One link that ties these two processes together is the dimorphic switch between yeast-like and filamentous forms. In model organisms such as the ascomycete yeast, Saccharomyces cerevisiae, and the basidiomycete smut, Ustilago maydis, membrane proteins that sense and transport ammonium have been implicated in this switch. The MEP proteins from yeast have been intensively studied, while those in other fungi, particularly pathogenic organisms, have been much less scrutinized.
The yeast S. cerevisiae can adopt several alternative developmental fates depending on the availability of specific nutrient sources. When nitrogen and fermentable carbon sources are both plentiful, haploid and diploid yeast cells reproduce by budding. When nitrogen is limiting but an abundant fermentable carbon source is present, diploid yeast cells undergo pseudohyphal differentiation (i.e., a dimorphic switch) to form filamentous colonies that forage for nutrients. These alternative fates allow this nonmotile organism to appropriately adapt to its surroundings. The signaling cascades by which yeast cells sense and respond to nutrients serve as models to understand how all cells sense and respond to the environment. Ammonium is a preferred nitrogen source for many fungal and bacterial species and for plants, as well. Moreover, the ability of microbes to sense and transport nitrogen is critical, not only for survival, but also as a prelude to a variety of developmental processes. These responses to nutrient availability are governed by both a mitogen-activated protein kinase (MAPK) cascade and the cAMP-dependent protein kinase pathway, with cross-talk between these parallel pathways at several important points; these cascades also control several aspects of pathogenic development discussed below. How nutrient signals in the extracellular environment control the activation of these globally conserved signaling cascades is a central question.
1. Nutrient sensing controls pathogenicity
Many fungal pathogens utilize a similar switch between a yeast-like form and a filamentous form as an integral part of their overall strategy of disease production. For example, the dimorphic transition from budding yeast to filamentous forms is crucial for infection for Candida albicans, and it is important for Cryptococcus neoformans. Similarly, in some plant pathogens, such as the maize pathogen Ustilago maydis, the dimorphic switch plays a critical role in both morphogenesis and pathogenicity. In fact, there is a high degree of conservation in the signaling cascades controlling development and virulence of these divergent ascomycete (S. cerevisiae, C. albicans) and basidiomycete (C. neoformans, U. maydis) yeasts. Based on these conserved pathways, studies in the model yeast S. cerevisiae have provided insight into fungal pathogenesis in humans and plants. In turn, studies in the pathogens will not only validate studies from yeast but also are expected to begin to identify universal and species specific components of ammonium sensing networks.

Several factors can cause cell differentiation in Dimorphic Fungi
Mating Pheromones
- Nutrients (low N)
- Host Cues
Nutrient sensing in fungal plant pathogens
As mentioned above, many of the conserved features of signaling are also found in fungal plant pathogens. The smut fungus U. maydis exists as haploid yeast-like sporidia, and mating of cells of opposite mating-type, accompanied by signals from host plants, leads to formation of infectious dikaryotic hyphae. U. maydis infection causes tumor formation on all maize shoot tissues, including tumor formation, with particular severity on the ears. Such damage can result in annual losses exceeding $200 million. For U. maydis mating occurs readily on rich media, but haploid or diploid sporidia grow filamentously on nitrogen starvation medium. The first genes encoding proteins involved in ammonium transport were cloned from yeast and Arabidopsis. As mentioned above, for S.cerevisiae, low nitrogen in the presence of an abundant fermentable carbon source leads diploid yeast cells to grow as pseudohyphae. The genes primarily responsible for both transport and sensing of available nitrogen include MEP1, MEP2, and MEP3, each of which encodes an integral membrane protein that transports ammonium ions or the toxic analog methylammonium. Of the three,Mep2 is a high-affinity permease, that acts as both an ammonium transporter and sensor. The yeast Mep2 protein controls the transition to pseudohyphal growth in response to ammonium limitation. mep2/mep2 mutant strains have no vegetative growth defect on nitrogen limiting medium, but in contrast to wild type cells are unable to undergo pseudohyphal differentiation. This defect can be suppressed by exogenous cAMP or by dominant activation of the PKA signaling pathway, supporting models in which Mep2 either functions upstream of PKA signaling, or the two pathways function in parallel. In U. maydis, we have identified two homologs of Mep2, called Ump1 and Ump2, and importantly Ump2 is required for filamentous growth on low nitrogen medium and can restore both growth and filamentous growth when heterologously expressed in S. cerevisiaemep1,2,3 mutant strains. Ump2 was identified as being more highly expressed under conditions of low ammonium. Thus, the S. cerevisiae Mep and U. maydis Ump proteins are physical and functional homologs of each other and function in both ammonium transport and ammonium sensing. The yeast Mep and fungal Ump proteins are part of a larger family of proteins that are conserved between bacteria, yeast, fungi, and humans.
2. Regulation of transporter function by phosphorylation
Several examples exist where proteins involved in transport are regulated by phosphorylation/dephosphorylation cycles. Our preliminary work examining the phosphorylation of the yeast Mep proteins indicates that a putative PKA site is present. Furthermore, the target residue is necessary for appropriate Mep function to control filamentation in both S. cerevisiae and U. maydis. Our ongoing studies aim to define the function of this phosphorylation, and will provide insight into conserved regulatory mechanisms that operate in these and other sensory receptors. We are also using split-ubiquitin and protein association assays to identify interactions between the Ump proteins and possible signaling components of U. maydis.
3. Additional Roles for Signaling Pathway Components
Proteins of the 14-3-3 and Rho-GTPase families are functionally conserved eukaryotic proteins that participate in many important cellular processes such as signal transduction, cell cycle regulation, malignant transformation, stress response, and apoptosis. However, the exact role(s) of these proteins in these processes is not entirely understood. Using the fungal maize pathogen, Ustilago maydis, we were able to demonstrate a functional connection between Pdc1 and Rho1, the U. maydis homologues of 14-3-3epsilon and Rho1, respectively. Our experiments suggest that Pdc1 regulates viability, cytokinesis, chromosome condensation, and vacuole formation. Similarly, U. maydis Rho1 is also involved in these three essential processes and exerts an additional function during mating and filamentation. Intriguingly, yeast two-hybrid and epistasis experiments suggest that both Pdc1 and Rho1 could be constituents of the same regulatory cascade(s) controlling cell growth and filamentation in U. maydis. Overexpression of rho1 ameliorated the defects of cells depleted for Pdc1. Furthermore, we found that another small G protein, Rac1, was a suppressor of lethality for both Pdc1 and Rho1. In addition, deletion of cla4, encoding a Rac1 effector kinase, could also rescue cells with Pdc1 depleted. Inferring from these data, we propose a model for Rho1 and Pdc1 functions in U. maydis, as follows. Rac1 sequesters Cla4 to the growing tip. Thus, Rho1 could interfere with Rac1 activities by preventing Rac1 from localizing at the growing pole. Other possibilities for how Rho1 could indirectly act as negative regulator of Rac1 are by sequestering Cdc24 (the RhoGEF) or Cla4 away from Rac1. Yeast two-hybrid analyses suggest possible interaction between Cdc24 and Rho1. As with Rho1, Cdc24-null cells are nonviable. In vivo, Rho1 and Rac1 could compete for Cdc24. On the other hand, Cla4 has been shown to localize at the bud-neck in S. cerevisiae. According to our GFP-Rho1 data, Rho1 also localizes (possibly with the help of Pdc1) at the bud-neck or in the septation area. Thus, it is a possibility that Rho1 is somehow involved with sequestration of Cla4 to the septation area and away from the growing tip and Rac1. In such cases, cells would then undergo budding instead of filamentation.
4. Control of Mitochondrial Inheritance
The mitochondrion is a critical organelle, whose importance far exceeds its common function as the energy-producing “powerhouse” of the cell, as it has been found to be involved in fundamental processes such as apoptosis, aging and metabolic homeostasis (Seo et al., 2010). Thus, appropriate inheritance of mitochondria is essential for growth and development of progeny. More importantly, such controls and the negative consequences of their disruption, have important ramifications for human health. Sexually reproducing eukaryotes have evolved similar mechanisms that allow the mitochondria of a single parent to be passed on to the offspring (uniparental inheritance or UPI). This pattern of UPI is nearly universal across eukaryotes, from isogamous protists with equal-sized gametes, to anisogamy, as seen in animals and plants with extreme gamete-size asymmetry. UPI may facilitate purifying selection against deleterious mutations, restrict intergenomic conflicts and limit mitochondrial heteroplasmy, the case where both parental types occur and are maintained in cells. On the other hand, cytoplasmic mixing reduces variation, impeding the efficacy of selection against defective organelles or selfish genetic elements (Razvilavicius et al. 2017).
The mating type contributing the majority of mitochondria to the next generation is “maternal”, while the other is “paternal”. Control of mitochondrial inheritance can similarly be maternal or paternal. In such cases, nuclear genes in one mating type control destruction of the mitochondria in the gametes of one particular mating type. Maternal control involves destruction of the partner’s mitochondria after fertilization, whereas in paternal control, nuclear genes in one mating type control destruction of its own mitochondria during gamete formation. Such controls may be found in isogamous organisms, but for multicellular organisms where there is gamete asymmetry, maternal control would amount to the targeting and elimination of mitochondria from sperm post-fertilization. In contrast, under paternal control, the exclusion or disabling of mitochondria occurs during spermatogenesis (before entering the oocyte). It is expected that paternally controlled organelle destruction is precluded because of the lack of long-term linkage between the paternal nuclear genotype and its own mitochondria, as the cytoplasm is exclusively maternally inherited (Razvilavicius et al. 2017). Such theoretical predictions typically explore isogamous systems, but should apply to multicellular organisms, since a paternal nuclear gene that causes the exclusion of sperm mitochondria cannot build up linkage with maternally inherited organelles, as this relationship is re-set every generation (Razvilavicius et al., 2017).
The pathogenic smut fungus Ustilago maydis (and its close relative Sporisorium reilianum) serves as an excellent model to study inheritance of mitochondria in higher eukaryotes (Fedler et al., 2009). The life cycle of this pathogen has been extensively characterized and its common host is Zea mays (maize or corn). The growth form of wild type U. maydis cells and pathogenicity on maize are inextricably linked because the filamentous, dikaryotic stage is the natural pathogenic cell type. The haploid strains of the fungus are saprophytic, budding cells that are easily cultured and manipulated in the laboratory. When two compatible U. maydis haploid cells are in close proximity, they produce fusion hyphae that grow towards each other and eventually fuse to form the infectious dikaryon; this cell type is an obligate biotroph that requires maize tissue for proliferation and development. Compatible haploid cells fuse and form a stable dikaryon only if they carry genes with different specificities at both the a and b mating-type loci. Cell fusion is controlled by the a mating-type locus, which has two alternative non-allelic forms (termed, a1 and a2 idiomorphs). The a locus encodes pheromones and pheromone receptors similar to those of yeast. Notably, U. maydis contains genes for the appropriate segregation of parental mitochondria during mating. The lga2/rga2 system controls a degradation-mediated UPI mechanism, in which all offspring will contain mitochondrial material from a single parent (a2). This mechanism is consistent with the several mechanisms of maternal mitochondrial inheritance seen in higher eukaryotes. Previous studies have elucidated the relationship between the lga2/rga2 system and the mating program of U. maydis. Pheromone stimulation upon the encounter between compatible partners confers low-level expression of the lga2/rga2 system in the a2 partner (Urban 1996). The protein product of rga2 has been linked to protection of a2 mtDNA, while the protein product of lga2 degrades unprotected mtDNA by a currently unknown mechanism that ultimately leads to selective mitophagy (Fedler et al., 2009). Heterozygosity at the multiallelic b locus is required for the production and maintenance of a stable filamentous dikaryon, and for pathogenicity (Brachmann et al., 2004). The b mating-type locus encodes two homeodomain containing proteins, bE and bW, that interact when produced from different alleles. The b heterodimer controls events after cell fusion required for the production of a stable filamentous dikaryon and regulates the transcription of a set of target genes that directly or indirectly control morphogenetic transitions and pathogenicity (Brachmann et al., 2001, 2004). After conjugation and subsequent cellular fusion of partners, the organism enters the pathogenic stage of its life cycle. The heterodimeric transcription factor bE/bW is upregulated during pathogenic development and is responsible for the upregulation of genes involved in host infection (Heimel et al., 2010). Interestingly, the bE/bW complex upregulates lga2 expression, leading to the degradation of unprotected a1 mtDNA (Fedler et al., 2009).
Sporisorium reilianum, another smut fungus, is a close relative of U. maydis, as evidenced by phylogenies based on ribosomal DNA sequencing as well as comparative genomics. They share a similar life cycle; nonetheless, mating in S. reilianum can occur between three parental strain types, a1, a2 and a3 . Interestingly, the a2 form in S. reilianum also houses orthologues of the lga2/rga2 mitochondrial inheritance system. As a result, offspring resulting from a cross between the a2 parent and either a1 or a3 are expected to be of the a2-mitotype. However, what occurs in a cross between the a1 and a3 strains? We propose that a cross between a1 and a3 partners will produce heteroplasmic offspring (a1/a3-mitotype). We predict that, given the importance of mitochondria in an organism’s development and the genetic conflict that heteroplasmy imposes, offspring of the a1/a3-mitotype will be less fit for survival. We are currently exploring these questions in collaboration with Dr. Michael Menze in the Biology Department here and with Dr. Jan Schirawski at the Matthias-Schleiden-Institut, in Jena, Germany.
Evolution of Antibiotic Resistance in Bacterial Populations
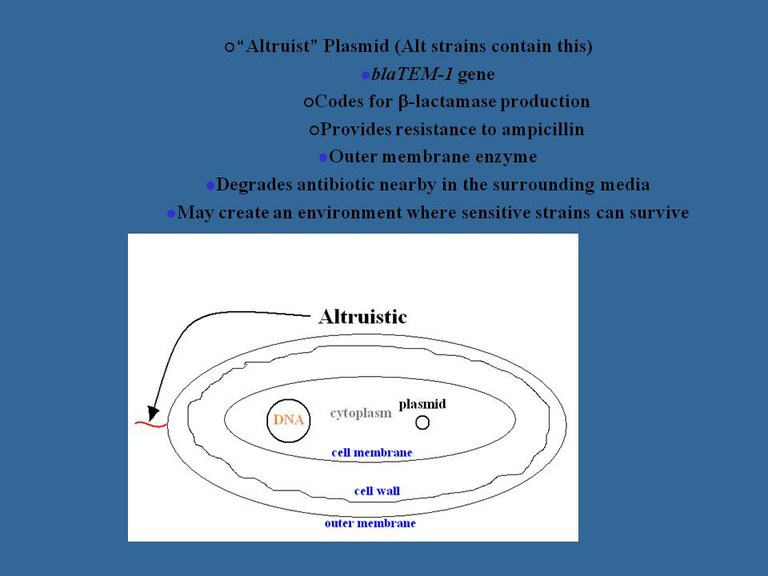
Microbial diversity is important in a variety of contexts, in particular having implications for infectious diseases, bioremediation, and environmental engineering. Examination of the mechanisms underlying such diversity and its evolution are important for providing a roadmap that can lead to better understanding of the above-mentioned and other, related, areas. Several examples are available where mutualism in microbial communities can lead to the maintenance of microbial diversity. Fewer examples have been presented where some individuals provide protection to others in a population without a concomitant benefit being returned to the protector. Protection of sensitive members of a population against antimicrobials by resistant genotypes has been observed in biofilms. In these cases, spatial proximity to the protector/producer was a prerequisite, or at least an important component, for survival by otherwise sensitive individuals. We have extended these studies to shaking liquid or planktonic populations in order to examine the dynamics of such unrewarded protection or altruism. We have developed a family of mathematical models that have examined antibiotic resistance in such systems (Dugatkin et al., 2003, 2005) and have generated preliminary data which support some aspects of these models (Dugatkin et al., 2004). Our results so far show that such altruists can provide frequency-dependent antibiotic resistance to other members in the population. Furthermore, frequency-dependent selection of the traits of these altruists promotes microbial diversity. We are currently conducting more detailed experiments designed to test our model's predictions, and will then use the results obtained to further refine the models. Specifically, we will use competitions between near-isogenic Escherichia coli strains that are either sensitive to ampicillin or are resistant, by virtue of a plasmid-encoded b-lactamase. Among those strains that are resistant, we compare the outcome for those strains that only protect themselves from ampicillin versus those strains that can also provide protection to others in their vicinity, i.e., by destroying ampicillin nearby in the medium. These analyses have also been extended to competition experiments with greater relevance for natural clinical or human settings. Specifically, we have examined whether ampicillin-resistant E. coli can protect sensitive Salmonella in their vicinity. Our results (Perlin et al., 2009) demonstrate clearly that non-pathogenic antibiotic-resistant bacteria may protect otherwise susceptible pathogens by a mechanism that does not involve gene transfer. Evolution of such transient survival mechanisms may thereby hinder therapeutic use of antibiotics, and should be considered in devising effective treatment strategies.

The Brandenberg Gate in Berlin. The program provides scientific and cultural opportunities. |
National Science Foundation International Research Experiences for Students (IRES)
Track I IRES Sites: Training with Smut Fungi in Germany
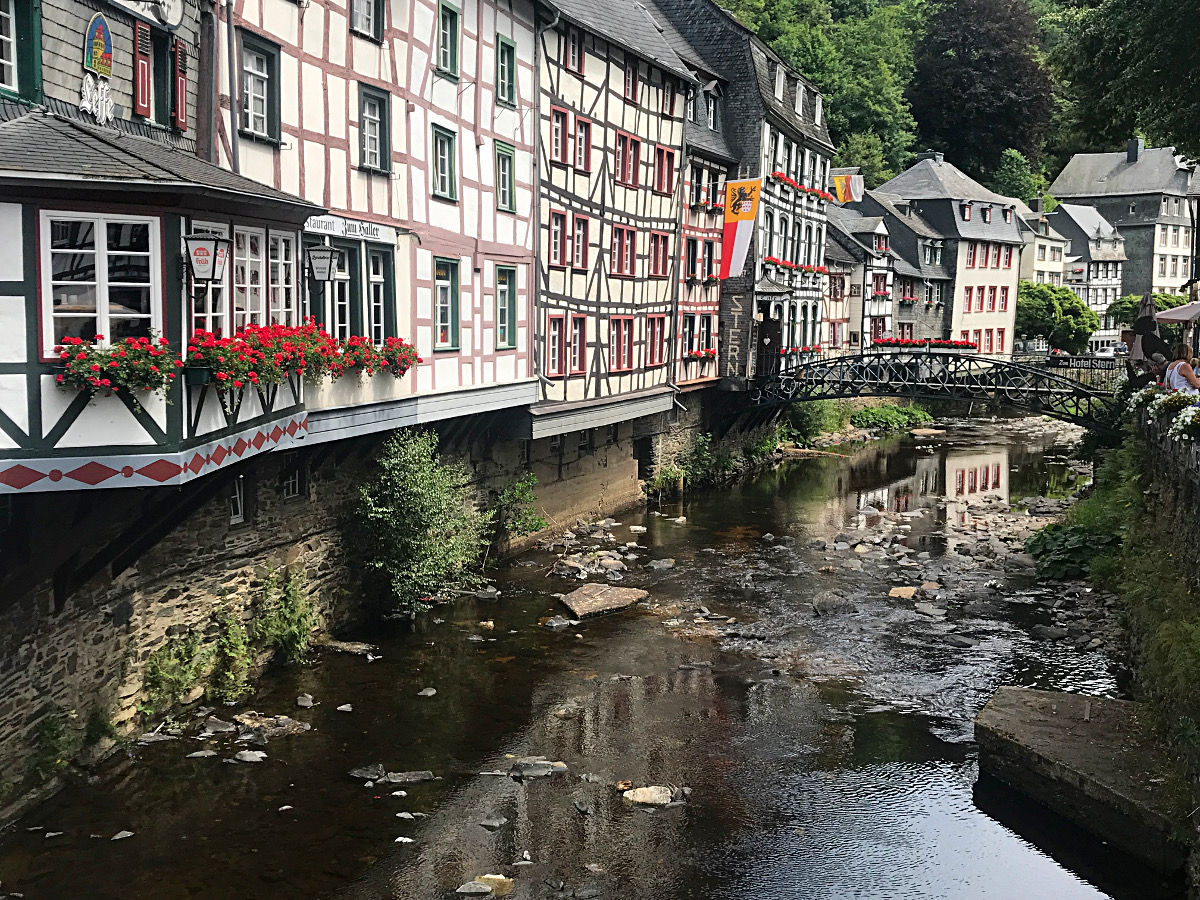
Through a three-year grant from the US National Science Foundation, our lab has developed an exchange program whereby graduate students and undergraduates from my lab visit the labs of research collaborators in Germany from 8-12 weeks each summer. Students accepted for these exchanges are trained and prepared during the normal academic year and then undergraduates are paired with a US graduate student to work on independent projects of mutual interest, to the students and to the labs involved in the collaborations. We endeavor to recruit members of groups traditionally under-represented in the sciences, both to expand opportunities for such student populations and to also provide all students chosen to expand their cultural horizons by being able to experience different ways to approach science and to view the world.
New landscapes and experiences await. | ||
Year 1 (May-September, 2019)
In Fall 2018, we recruited 3 Doctoral (1 Hispanic/Latino and 2 White non-Hispanic students) and 3 undergraduate students (1 African-American female, 1 White non-Hispanic female, and 1 White non-Hispanic male) as candidates for this project. This group was trained in basic molecular biological techniques and in some aspects of species-specific techniques prior to travel. In general, undergrads were paired with their graduate student mentors, both for training purposes and so that they would become familiar with each other on a working basis prior to their visit to Germany. In addition, all students received formal or informal training in elementary German language.
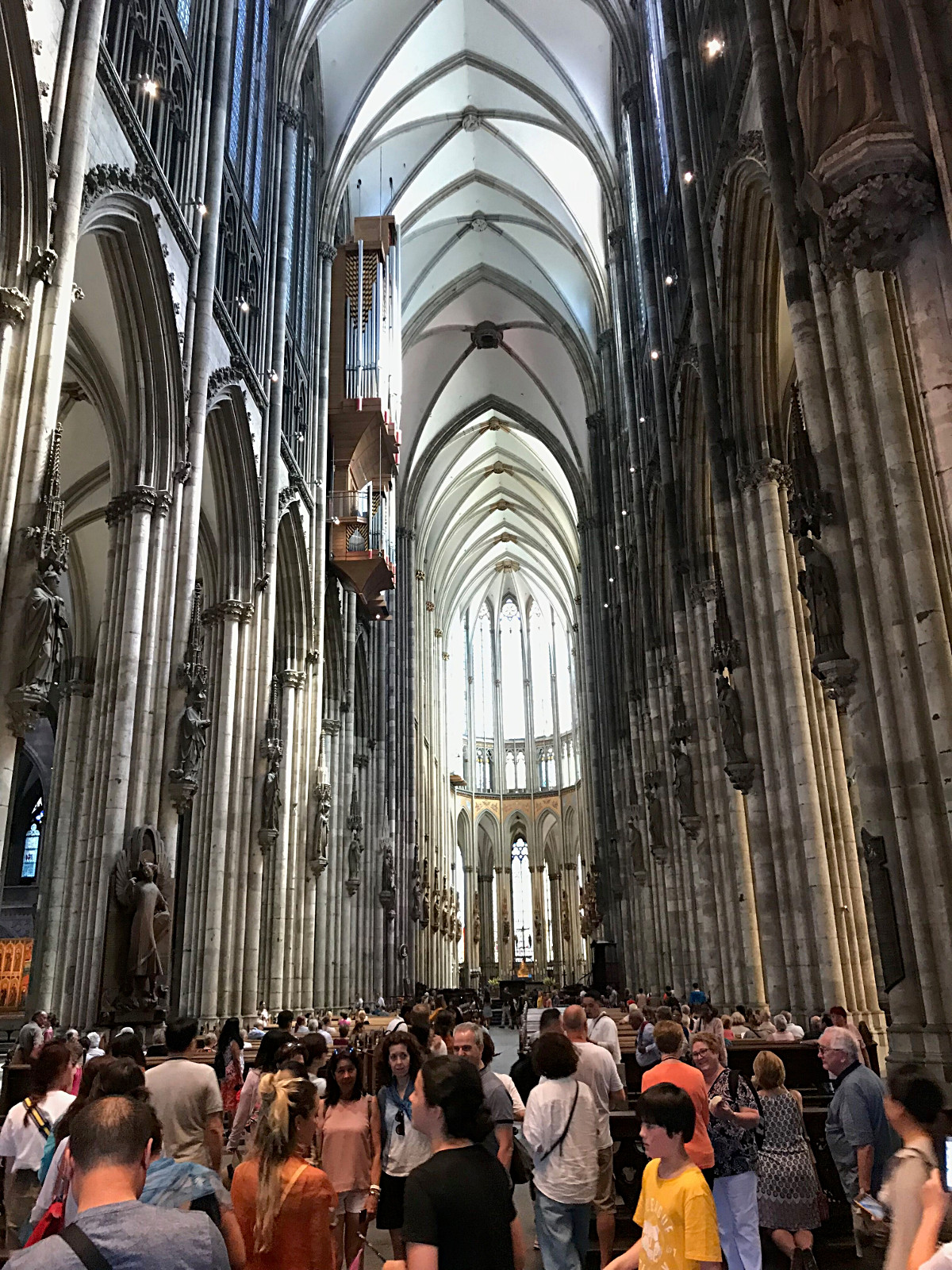
Joining the crowd in Koln Cathedral. |
On May 15, 2019, students began their research visits to their respective labs: 2 graduate student/undergrad pairs went to Jena, Germany to the lab of Prof. Dr. Jan Schirawski, Matthias-Schleiden-Institut. These students are working on questions relating to Sporisorium relianum; specifically, they are investigating fungal proteins that manipulate host plants or are investigating mitochondrial inheritance and mitochondrial function in this organism. One graduate student/undergraduate pair joined the lab of Dr. Dominik Begerow at the RWTH University of Bochum. There they are investigating ways of genetically manipulating M. violaceum strains, conducting large-scale greenhouse experiments to investigate relative pathogenicity of strains, and learning microscopy techniques to follow the infection process.
|
||
|
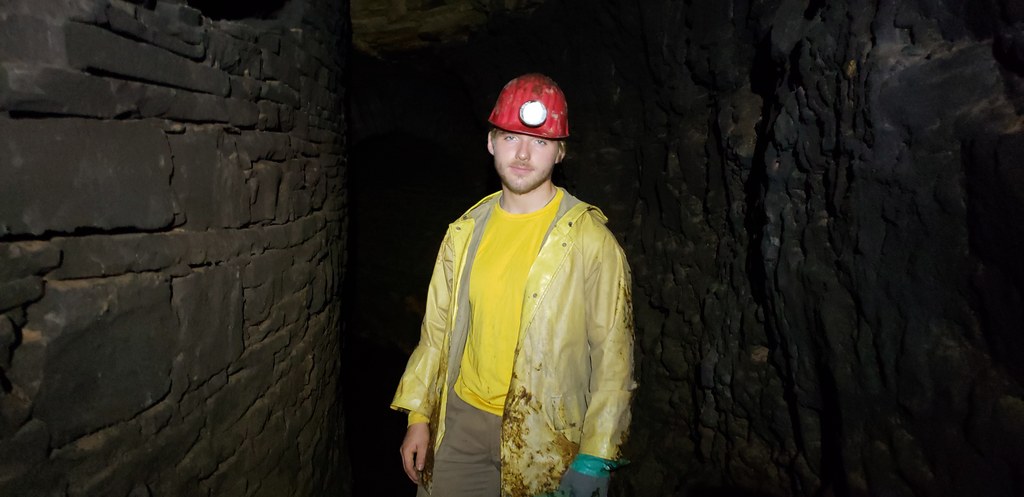
In addition to research trips to dark places, participating students can engage in research brainstorming sessions and, possibly, other traditional forms of inspiration. |
|
Major Research Goals:
Bochum Group:
Graduate Student (William Beckerson): Develop and compare methods for implementing CRISPR Cas to disrupt genes in the fungus, Microbotryum lychnidis-dioicae. This involves alternative methods for introduction of the Cas9 enzyme into cells and improved methods of fungal transformation.
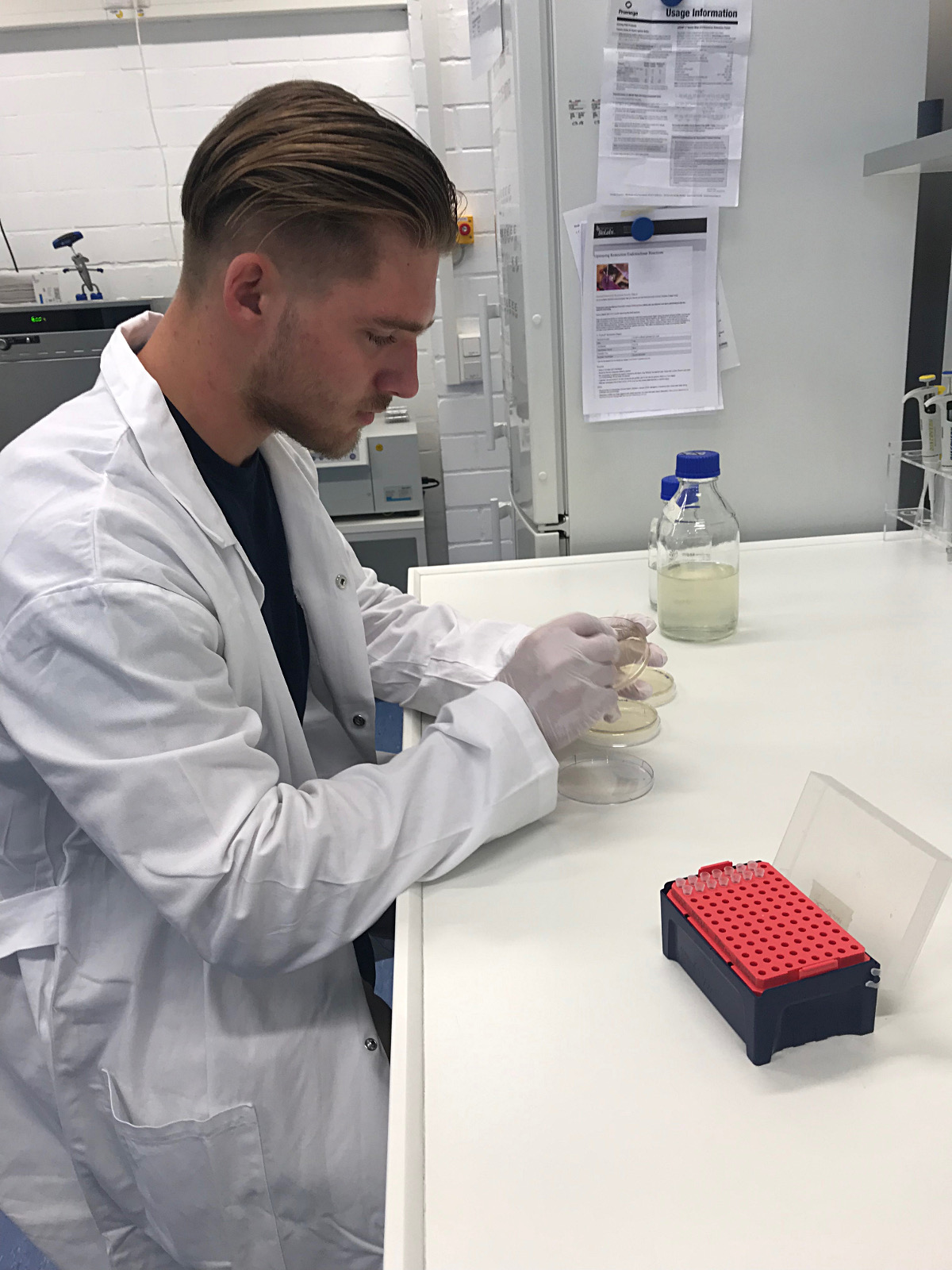
Phillip Sullivan in the greenhouse. |
Undergraduate student (Phillip Sullivan): Analyze the relative virulence of wild type infection by Microbotryum species on their natural hosts compared with those of M. lychnidis-dioicae modified to express a putative effector from the sister species, M. silenes-dioicae.
Jena Group:
Graduate Student (J. Ham): Express an effector from M. lychnidis-dioicae in Sporisorium relianum. Test such recombinant strains on the maize host to see if they elicit symptoms already observed in the Microbotryum hosts, e.g., early flowering.
Undergraduate Student (Kiara Smith): Express an effector from S. reilianum in M. lychnidis-dioicae and examine whether these strains lead to suppression of apical dominance, as observed in the natural interaction.
Graduate Student (Hector Mendoza): Conduct infection studies of S. reilianum on maize to compare relative virulence of different combinations of mated strains, in order to interrogate the roles of mating type on this process. Also, examine mitochondrial inheritance after such infections.
Undergraduate Student (Caroline Culver): Generate a pair of S. reilianum strains with their alternative oxidase (AOX) genes disrupted. Test these for susceptibility to stresses and relative virulence.
The Major Training Goals of this project were to:
1) Recruit both Doctoral student and Undergraduate student participants, preferably from URM groups.
2) Prepare them ahead of their travel for an optimal experience in the German labs participating in this project.
3) Give them training specifically in work with smut fungi (i.e., Sporisorium reilianum, Ustilago maydis, Microbotryum violaceum spp.)
4) Provide them with specialized training in microscopy.
5) Provide them with intellectual and cultural experiences in the host country.
Questions about this program and possibilities for joining this group may be directed to Dr. Perlin at anthersmut@louisville.edu
Phillip Sullivan at the bench |
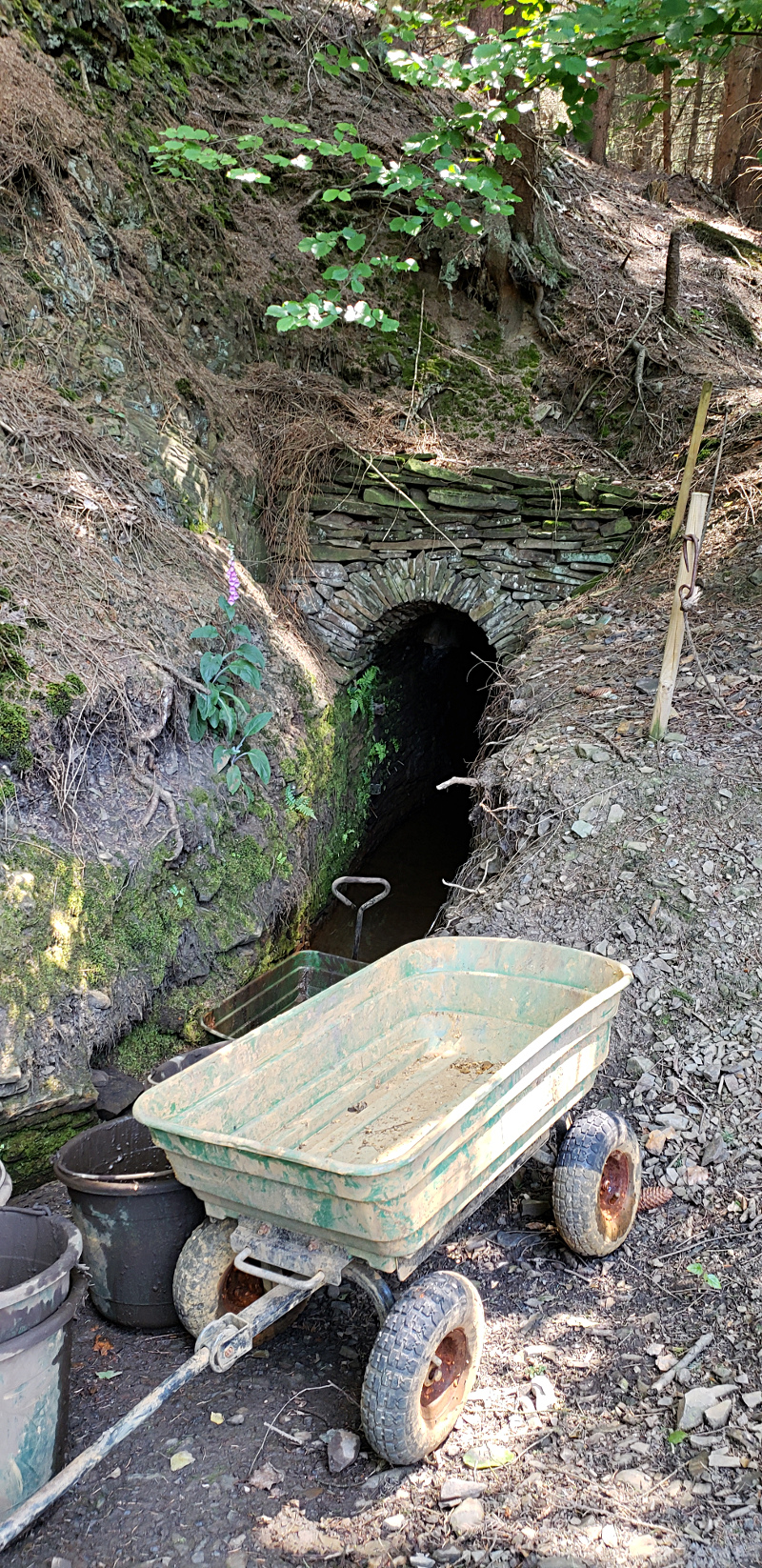
Local cultural experience: Community mine restoration project |
A display of mustards |
Current Students in Dr. Perlin’s lab
William Beckerson at the bench. |
William Beckerson, recent PhD. Examined fungal effectors across the Microbotryumviolaceum species complex; developed CRISPR/Cas9 technology to disrupt genes for putative effector genes so as to functionally test their roles in manipulation of host plants. Moving on to a Post-doctoral position exploring "Zombie ant" fungi at the University of Central Florida, with Dr. Charissa de Bekker.
Sunita Khanal, recent PhD. Characterized genes involved in nitrogen assimilation in U. maydis. Moving on to a Post-doctoral position at the University of Maryland Cardiology Division, developing CRISPR/Cas9 to study myosin phosphatase in hypertension and other vascular disorders with Dr. Steven Fisher.
Hector Mendoza, graduate student. Examining mechanisms for control of mitochondrial inheritance and fitness costs associated with its evolution in S. reilianum and U. maydis.
Ming-Chang "Nelson" Tsai, graduate student. Studying roles of gene expression for various target genes in M. violaceum.
Joseph P. Ham, graduate student. Studying the roles of fungal effectors and whether they can alter host preference or host manipulation across species boundaries.
Roxanne Leiter, graduate student. Investigating the roles of fungal effectors in establishing whether a fungus will be a “specialist” and only be able to infect one or a few host species or a “generalist” able to infect a broad range of host species.
Shikhi Baruri, graduate student.
Otniel “Alex” Nava-Mercado, undergraduate. Working on nitrogen assimilation in U. maydis.
Caroline Culver, undergraduate. Investigated role of alternative oxidase function in U. maydis and S. reilianum. Currently enrolled in University of Louisville medical school.
Grace Long, undergraduate. Examining possible host interactors with fungal effectors.
Phillip Sullivan, undergraduate. Exploring the role of host-specific effectors on host range.
Luke Schroeder, undergraduate. Working on nitrogen assimilation in U. maydis.
RECENT FORMER STUDENTS OR SOON-TO-BE GRADUATES
Lalu Villa Krishna Pillai, PhD. Identified a role for a PTEN orthologue in U. maydis in teliospore development and germination, as well as in pathogenicity. Currently Research Regulatory Specialist Norton Healthcare.
Su San Toh, PhD (deceased). Examined gene expression in plants infected with M. violaceum. Participated in the first genome and transcriptome work for these fungal species. Helped develop the first reproducible genetic transformation system for these fungi. Author, co-author, or collaborator on over 11 publications about this system.
Michael Cooper, PhD. Exploring nitrogen metabolism in U. maydis. Currently working at Murray State University.
R. Margaret Wallen, PhD. Examined the interaction between ammonium transport and mating in U. maydis. Currently, conducting research on endosomes in communication between different cell types; also, teaching biology at the college level.
Swathi Kuppireddy, PhD. Characterized fungal effectors for M. violaceum and examined the role of genes in controlling their host plants.
Hannah Mianzo, currently, Genetic Counselor.
Ben Lovely, PhD. Examined signaling via Hsl proteins in U. maydis and their connections to other signaling pathways. Currently, faculty, Department of Biochemistry and Molecular Biology, University of Louisville Medical School.
Charu Agarwal, PhD. Co-mentored with Dr. David Schultz, Department of Biology. Charu analyzed cAMP turnover in U. maydis via a genetic and biochemical analysis of cyclic phosphodiesterases of the fungus.
Jinny Paul, PhD. Characterization of ammonium transporter homologues of several fungi, with emphasis on those from U. maydis and M. violaceum.
Cau Pham, PhD. Exploring the roles of 14-3-3 and Rho proteins in cytokinesis, cell polarity, morphology, and pathogenesis for U. maydis. Currently a Research Associate at the Centers for Disease Control.
Zhanyang Yu, PhD. Isolation and characterization of Rho1 homologue in U. maydis and identification of interacting proteins and pathways.
Kiara Smith, undergraduate, Honors student. Investigated role of nitrite reductase in U. maydis. Currently enrolled in University of Louisville medical school.
Jack Desmarais, undergraduate, Honors student. Investigated role of PTEN orthologue in U.maydis. Currently enrolled in University of Louisville medical school.
Madison Furnish, undergraduate. Investigated roles of mating-type on gene-specific function in U. maydis. Currently in PhD program on Pharmacology at the University of Colorado, Denver.
Jared Andrews, undergraduate, Honors student. Characterizing the roles of sugar transporters in cell morphology and mating in M. violaceum. Currently in PhD program at Washington University, St. Louis.
Dominique Razeeq, undergraduate. Characterizing the roles of secretory lipases in cell morphology and mating in M. violaceum. Currently in medical residency, after graduating University of Louisville medical school in 2019.
Trisha H. Patel, undergraduate. Member of the team identifying interacting proteins with signaling components from U. maydis that may affect nitrogen metabolism.
Ann-Claude Rakotoniaina, undergraduate. Member of the team examining gene expression of M. violaceum during plant infection.
Tia Alton, undergraduate, Honors Student. Member of the team working on competition experiments with bacteria. Currently medical physician.
Evan Raff, undergraduate, Honors Student. Member of the team identifying interacting proteins with signaling components from U. maydis. Currently medical physician.
Alexander Bajorek, undergraduate, Honors Student. Member of the team working on competition experiments with bacteria. Currently medical physician.
Courtney McKenzie, undergraduate, Honors Student. Member of the team working on competition experiments with bacteria. Currently medical physician.
Cayse Powell, undergraduate, Honors Student. Member of the team working on competition experiments with bacteria. Currently medical physician.
Himati Patel, undergraduate, Honors Student. Member of the team working on competition experiments with bacteria. Currently pharmacist.
Lacey Hazel, undergraduate. Work on developing a gene disruption system for M. violaceum. Currently applying to graduate programs in neurobiology.
Greg Shaw, undergraduate, Honors Student. Member of the team identifying interacting proteins with signaling components from U. maydis. Currently medical physician.
Trisha Patel, undergraduate, Honors Student. Member of the team identifying interacting proteins with signaling components from U. maydis. Currently medical physician.
Anna Hellman, undergraduate, Honors Student. Member of the team identifying interacting proteins with signaling components from U. maydis. Currently in Pediatric Neurology residency, University of Louisville.